Data Reduction
Data reduction can be done on the ASTE computers at the Site, San Pedro, and Mitaka, e.g., aste-1s, aste-at1, aste-mt1. You can also reduce the data on common-use computers at Nobeyama (not aste-w). If you have a Linux PC, you can use the Export version of the reduction software.
Copy Raw Data
Raw data are created in a directory /cosmos3/aste/raw/otfdata/GROUP/PROJECT/otfdata/raw at the ASTE site. The raw data file is named as TableName.ObjectName.ObservationDate(YYYYMMDDhhmmss).NumberOfSpectrometers.N (e.g., klotf.OriKL.20050131204055.4.N).
Raw data should be placed in a directory /home/GROUP/PROJECT/otfdata/raw/ :
- If you reduce the data at the site
- Copy raw data from /cosmos3/aste/raw/otfdata/GROUP/PROJECT/otfdata/raw/ to /home/GROUP/PROJECT/otfdata/raw/ :
- cp /cosmos3/aste/raw/otfdata/GROUP/PROJECT/otfdata/raw/RawDataFile /home/GROUP/PROJECT/otfdata/raw/
- Oherwise, make a symbolic link to /cosmos3/aste/raw/otfdata/GROUP/PROJECT/otfdata/raw as /home/GROUP/PROJECT/otfdata/raw :
- ln -s /cosmos3/aste/raw/otfdata/GROUP/PROJECT/otfdata/raw /home/GROUP/PROJECT/otfdata/raw
- If you reduce the data at San Pedro
- Copy raw data in /home/GROUP/PROJECT/otfdata/raw/ :
- cp RawDataFile /home/GROUP/PROJECT/otfdata/raw/
- If you reduce the data at NRO
- Copy raw data in /home/GROUP/PROJECT/otfdata/raw/ :
- cp RawDataFile /home/GROUP/PROJECT/otfdata/raw/
In all cases, do not modify the contents of the directory /cosmos3/aste/raw/otfdata/GROUP/PROJECT/otfdata/raw/ .
Startup the Reduction Software
- If you reduce the data at the site or San Pedro
- Run the startup script:
- /home/asteotf/pro/nostar
- If you reduce the data in NRO
- Run the startup script with an option "-aste": i.e.,
- /home0/otf/IDL/nostar -aste
- It is strongly recommended to run the script on a CPU server (mpu1u, mpu2u, mpu4u, mpu5u, or mpu6u).
Input Group and Project and push "Start", then the Button Panel appears.
Fig. 4-1: Startup window of the reduction software.

Fig. 4-2: the Button Panel of the software.
Split
The raw data contains spectra from all the spectrometers used (A1--A4 for ACG, or X1--X12 for WHSF). At first, the raw data should be split into "Split raw data" files, each file contains spectra from a single spectrometer. The "Split raw data" is written in a directory /home/GROUP/PROJECT/otfdata/split . The raw data file is named as TableName.ObjectName.ObservationDate.NumberOfSpectrometers.N, and the split raw data is TableName.ObjectName.ObservationDate.NumberOfSpectrometers.N.NameOfSpectrometer (e.g., klotf.OriKL.20050131204055.4.N.A01).
By clicking "Split" button in the Button Panel, GUI for "Split" like Fig. 4-3 appears. Add raw data file(s) to be split using "Add File(s)" or "Wildcard" button. Click "Execute", then the split raw data files are written in /home/GROUP/PROJECT/otfdata/split .
- "Add File(s)": select file(s) via a file selection dialog box.
To select multiple files, control-click or shift-click on file names in the dialog.
#Note: the program will hang up if you select too many (typically several hundred) files at once (it seems that an IDL built-in function DIALOG_PICKFILE has an limit, 32768 characters, on total length of full-paths of selected files). - "Wildcard": select file(s) using wildcard characters: e.g., "/bin/ls /home/a0011hg/proj0/otfdata/raw/*20070416*" (commands other than ls, such as find, also can be used).
- "Remove"/"Remove All": remove a file or all files from the list.
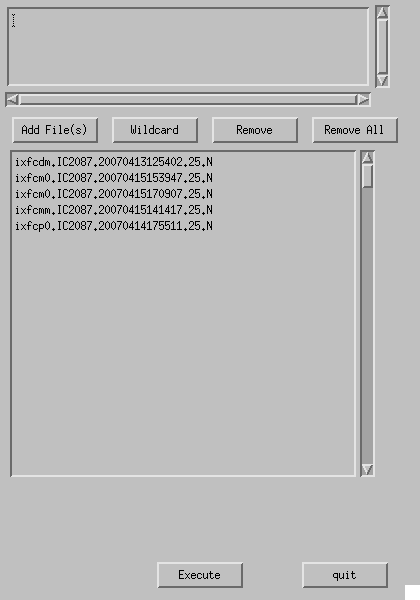
Fig. 4-3: Split.
Scale
If you want to scale the data, use a task "Scaling". This task should be used for split raw data files (no matter whether baselines have been subtracted or not). The values of scaling factors are stored in a header item "MLTSCF0". You can ascertain them using a task "Show Header".
By clicking "Scaling" button in the Button Panel, a window like Fig. 4-4 appears.
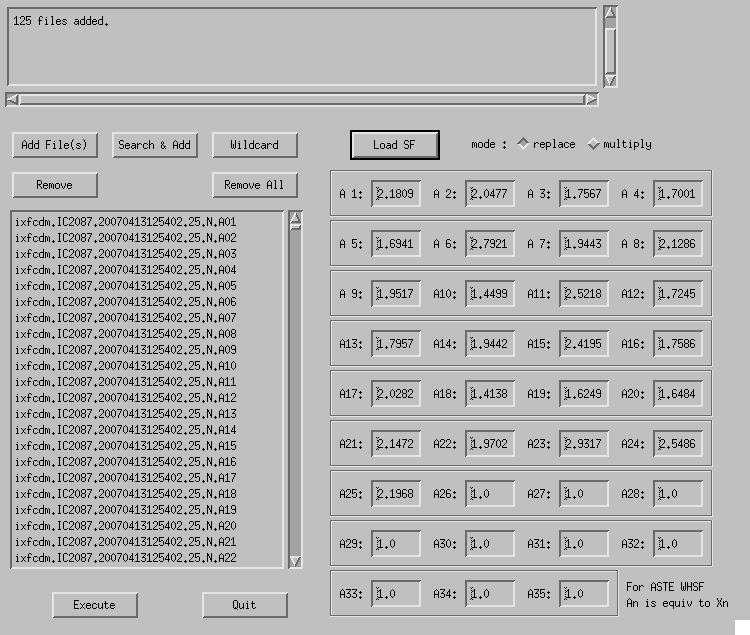
Fig. 4-4: Scaling.
Select split raw data file(s) by clicking "Add File(s)" or "Wildcard" button. Click "Search & Add" to automatically add data taken with other spectrometers. Input scaling factors for spectrometers "A1"--"A4" and click "Execute" then the factors are applied to the data.
- Scaling factors can be loaded from a text file using "Load SF". A set of name of a spectrometer An and corresponding scaling factor should be written in each line (a sample of scaling factor file). The names of spectrometers can be omitted: in that case, it is assumed that the values for A1, A2, ... are written in order.
- When "mode" is "replace", MLTSCF0 is overwritten with the given values. On the other hand, if "multiply" is selected, the given values are multiplied by the currently-set MLTSCF0 values.
Base
Fit and subtract spectral baselines from the split raw data. The file after baseline subtraction is named as TableName.ObjectName.ObservationDate.NumberOfSpectrometers.NameOfSpectrometer.base (e.g., klotf.OriKL.20050131204055.4.A01.base). The original split raw data file is deleted. Split raw data, for which baselines are not subtracted, are skipped in "Make Map".
Use "Baseline" or "Base (Batch)" in the Button Panel.
Baseline
File selection dialog (Fig. 4-5) appears by clicking "Baseline" button in the Button Panel. Click "File" to pick a split raw data file, and click "Load" to open GUI window (Fig. 4-6).
Fig. 4-5: File selection dialog for Baseline.
The data are displayed in the right-hand side of the window, while parameters are set in the left-hand side. Data display and baseline fitting are done for each scan row. The upper panel of the data display is an averaged line profile of the scan row. The lower panel shows a kind of position-velocity diagram: horizontal axis shows the velocity (or channel, frequency), and vertical axis represents the spectrum number.
- Specify the function form using "fitting function" and "order/wave"
- Input "fitting range". Up to 4 baseline ranges can be specified. Ranges are input by clicking on the data display, or manual input. When you click "clear" ranges are cleared.
- Specify unit and channel binning width using "x-axis mode" and "ch binning".
- Specify reference (rest) frequency using "Ref freq".
- Specify velocity- and intensity-axis range using "xrange" and "yrange". "yrange" also means the gray scale intensity range of the lower panel.
- By clicking "Exec", baseline subtraction is done for the currently-displayed scan row.
- "Next" and "Prev" are used to show the next and previous scan rows respectively.
- By clicking "Till Last SP", baseline subtraction is done for all the scan rows from the currently-displayed one to the last one.
- "Revert" reverts the file to the original state.
- The task finishes when "Save & Quit" is clicked.
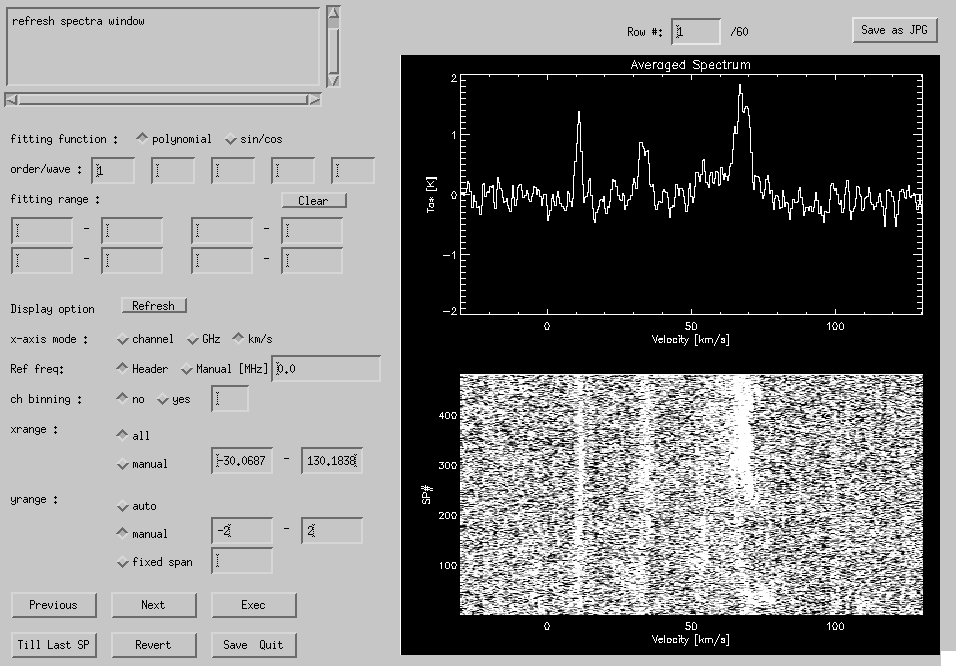
Fig. 4-6: Baseline GUI.
Base (Batch)
A window like Fig. 4-7 appears by clicking "Base (Batch)" button on the Button Panel. Add split raw data file(s) to the file list using "Add File(s)", "Search & Add" or "Wildcard". Specify the type and parameters (polynomial order or sinusoidal wave number) of baseline-fitting function in "function type", range and unit in "fitting range" and the fit is executed when "Exec" is clicked.
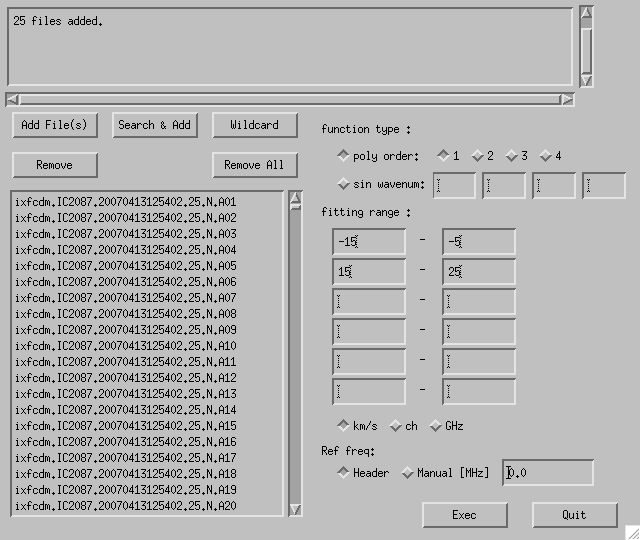
Fig. 4-7: Base (Batch) GUI.
Flag
(If necessary) use "Flag" to flag out bad data. The flagged data are not used in "Make Map" (flagged data are not removed, but only flagged).
File selection and option setting window (Fig. 4-8) appears when you click "Flag" on the button panel. Select a split raw data file (the data taken with other spectrometers will be simulataneously processed). In this task time-averaged spectra are shown (like the QLOOK). The time span for averaging is specified in "Time averaging" option. For "Divide each scan into", each scan rows are divided into the specified number. For "Number of spectra" the data are binned by the specified numbers from the beginning of the data.
When you click "Load" the program begins to make temporary files (with extension ".sp"). Please wait for a while. If previously-made ".sp" files are found, the program uses them instead (therefore ".sp" files are not erased when quitting the task).
Fig. 4-8: File selection dialog for Flag.
GUI window (Fig. 4-9) will appear when all the ".sp" files are made. Spectra are shown in the right-hand side of the window. "Cross" marks are drawn on flagged spectra (when only a portion of the data within the averaging span is flagged, a slanting line is drawn instead).
- Specify unit and channel binning width using "x-axis mode" and "x smoothing".
- Specify reference (rest) frequency using "Ref freq".
- Specify velocity- and intensity-axis range using "xrange" and "yrange".
- "Prev" and "Next" are used to show the previous and the next page respectively.
- By clicking "Exec", flag information is written into the data files.
- "Flag All CH" and "Unflag All CH" turn flags ON/OFF in the currently-displayed page for all the spectrometer.
- "Quit" to quit. If some of the flag information is not saved yet, the program confirms whether "Force quit" or "Save & quit".
- "Flag All Data" and "Unflag All Data" turn flags ON/OFF for all the data in all the spectrometer.
- Currently-displayed page number is shown in "Page #". Rewrite the number and hit Return to jump to a desired page.
Clicking on a spectrum toggles the flag. The following keyboard shortcuts are available when the mouse cursor is on the spectra:
- "b" or"p": previous page
- "f" or "n":next page
- "q": quit
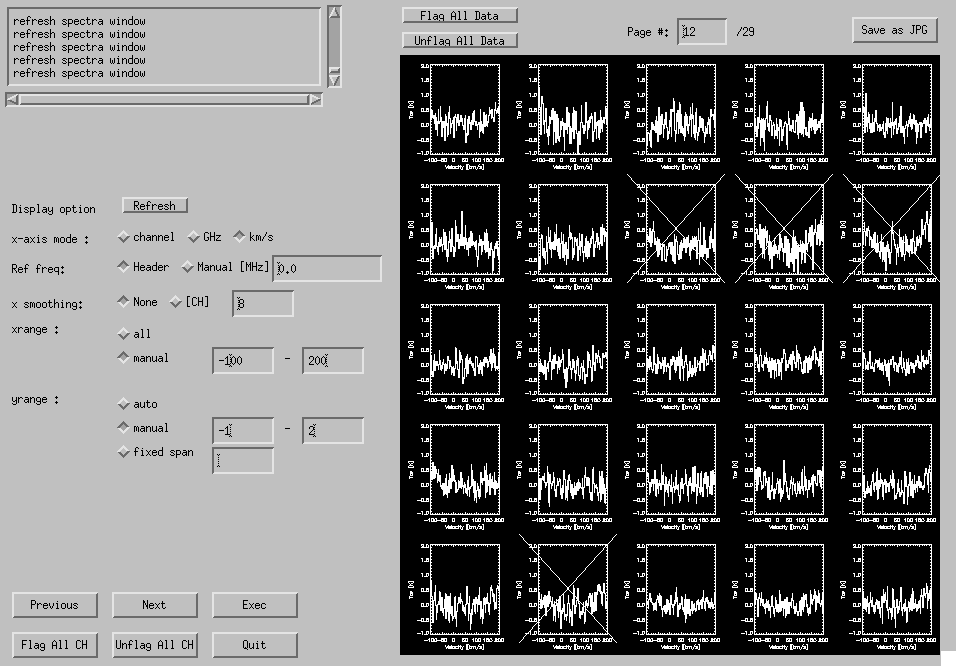
Fig. 4-9: Flag GUI.
Map
Finally a map (FITS cube) is created from baseline-subtracted split raw data files using convolution and regridding (see Introduction).
A file/directory selection window (Fig. 4-10) appears by clicking "Make Map" on the Button Panel. Use "Add File/Dir" or "Wildcard" to add split raw data files, or directories in which split raw data files exist (we recommend to use directories. To pick a directory on the file selection dialog box, move into the directory, and click "Open" button without selecting any file). Specify the coordinate system with "Position map type", and "Load" opens a GUI window (Fig. 4-11).
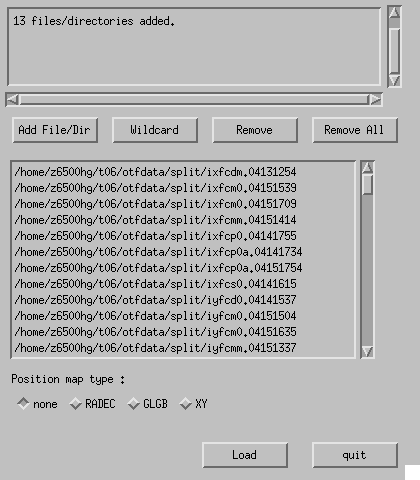
Fig. 4-10: File/directory selection window of "Make Map" task.
- Specify coordinates for the position map and mapping range using "Coordinates".
- Input the bottom-left corner (BLC) and the top-right corner (TRC) in "Mapping Range". Click on the position map, or manual input.
- Set grid spacing or number along the spatial axes using "Spatial Grid"
- Specify velocity (frequency) axis type using "Freq axis"
- Specify velocity range to be mapped using "Range".
- Set grid spacing or number along the velocity/frequency axis using "F/V Grid".
- Specify reference (rest) frequency using "Ref freq".
- Position map can be zoomed in and out. At first click "Zoom", and click BLC and TRC. "Unzoom" reverts to the original (default) state.
- Specify a convolution function and its parameters.
- Using a button "Load .his", parameters can be loaded from a history file (with an extension of ".his").
- Specify output FITS file name, and click "Exec". It will take time.
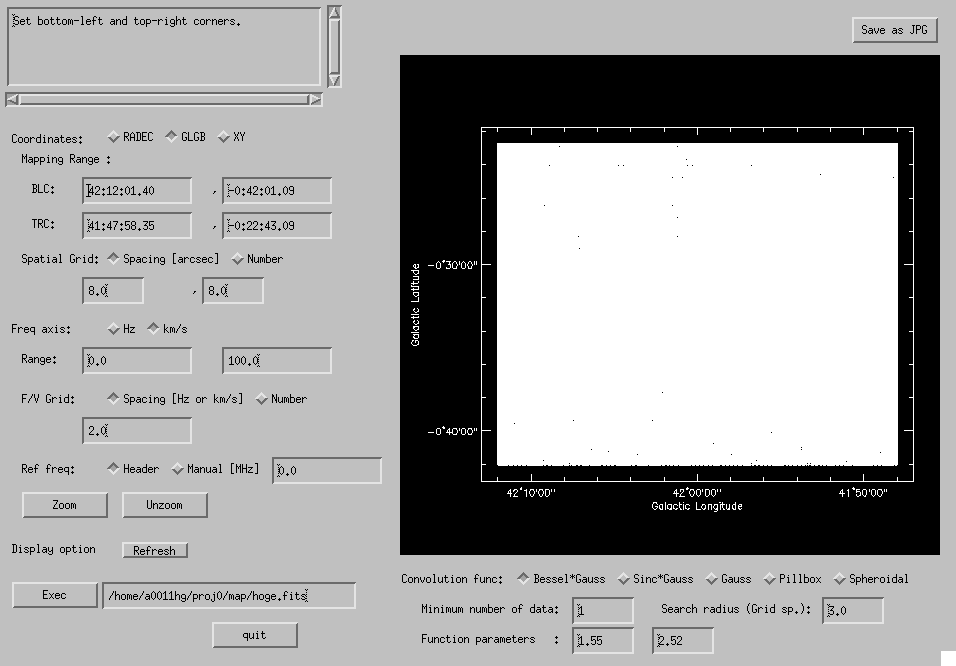
Fig. 4-11: Make Map GUI window.
When you execute "Make Map", in addition to output FITS cube hoge.fits, the following 3 files are created: a temporary file hoge.fits.grid, a history file hoge.fits.his, and a 2-D FITS hoge.fits.rms.fits in which rms value of each grid point of the map is written. Attention: rms values written in hoge.fits.rms.fits are rough estimates (smaller values than actual rms tend to be written), thus you have to judge the data quality using the output FITS cube itself. You may delete the ".grid" file (it's huge).
Basket-Weave
Two FITS images (cubes) are combined using the algorithm described by Emerson & Gräve (1988) (Fourier-transform the maps, mask the scanning noise in Fourier domain for each map, weighted-mean them, and inverse Fourier-transform) in order to remove the "scanning effect". The two input FITS files must have the same coordinate axes (i.e., the same NAXISi, CRVALi, CRPIXi, CDELTi, and CROTAi).
By clicking "Basket-Weave" in the Button Panel, a window like Fig. 4-12 appears.
- For both "Image 1" and "Image 2",
- Specify input FITS file name using "File" button.
- Set "Scanning direction (deg)". Set to 90 if the FITS image is made from X-scans; set to 0 if the FITS image is made from Y-scans.
- Set "R.M.S. noise level" of the maps. Inverse-square of the values are used as weights.
- Pixels with poor data quality can be substituted by BLANK values using "Clip"
- Set minimum and maximum intensity of valid data in "intensity". Pixels whose intensity is out of this range becomes BLANK.
- Set minimum value of RMS of the map in "RMS value". The RMS map, hoge.fits.rms.fits, is used to estimate the RMS value of each pixel.
- In "Option" the width of mask function applied to Fourier-transformed maps can be specified.
- In "Preview" fields, specify intensity minimum, maximum, and number of Z plane (velocity or frequency) to be previewed. Press "Show" to display the preview. In the preview area, "before & after" maps and their Fourier-transformed images are shown.
- Specify output FITS file name, and then click "Exec".

Fig. 4-12: Basket-Weave GUI window.
Command-Line Reduction
If you cannot open GUI windows through the network, command-line reduction can be made as follows. When you reduce the data in the Site, take care not to interfere the running observation. Commands are installed in /home/asteotf/bin/ (the site and San Pedro); /home0/otf/bin/ (NRO). In NRO, it is strongly recommended to run the commands on a CPU server (mpu1u, mpu2u, mpu4u, mpu5u, or mpu6u).
Split
Type
otf_split RawData
to execute Split. Where the raw data exist is described in "Copy Raw Data" section. The split raw data are written in a directory defined by an environment variable OTF_BASE_DIRO. If it is undefined, /home/GROUP/PROJ/otfdata/split/ is used. An error occurs if the destination directory does not exist, thus you should mkdir it in advance.
(e.g.) otf_split /home/a0011hg/proj0/otfdata/raw/klotf.OriKL.20050131204055.4.N
Scale
Type
otf_scale SplitRawData ScalingFactor Mode
to execute scaling. Mode is "r" or "m". When it is set to "r", the header item "MLTSCF0" is overwritten by the given scaling factor. When "m", the given factor is multiplied to the currently-set "MLTSCF0"
(e.g.) otf_scale /home/a0011hg/proj0/otfdata/split/klotf.OriKL.20050131204055.4.N.A01 1.42 r
Base
Type
otf_base SplitRawData StartSpNumber EndSpNumber FunctionType Range1StartChannel Range1EndChannel Range2StartChannel Range2EndChannel Range3StartChannel Range3EndChannel Range4StartChannel Range4EndChannel Range5StartChannel Range5EndChannel Range6StartChannel Range6EndChannel OrderOfPolynomial_or_WaveNumberOfSinusoid
to execute baseline. FunctionType is "polynomial" or "sincos". For "sincos", multiple wave numbers can be specified (as the last arguments) separated by spaces. Baseline ranges should be specified in channel, not velocity or frequency. The output file is written in a directory defined by an environment variable OTF_BASE_DIRO. Type
setenv OTF_BASE_DIRO /home/GROUP/PROJ/otfdata/split
in advance, otherwise you will meet an error.
(e.g.) otf_base /home/a0011hg/proj0/otfdata/split/klotf.OriKL.20050131204055.4.N.A01
1 9000 polynomial 150 250 750 850 0 0 0 0 0 0 0 0 1
(e.g.) otf_base /home/a0011hg/proj0/otfdata/split/klotf.OriKL.20050131204055.4.N.A01
1 9000 sincos 150 250 750 850 0 0 0 0 0 0 0 0 2.3 3.1 4.2
Map
Type
otf_map CoordinateSystem Projection ProjectCenterPositionX ProjectCenterPositionY BLCPositionX BLCPositionY TRCPositionX TRCPositionY ZaxisType ZaxisStartPoint ZaxisEndPoint XaxisGridNumber YaxisGridNumber XaxisGridSpacing YaxisGridSpacing ZaxisGridNumber ZaxisGridSpacing SearchingRadius MinumumNumberOfData ConvolutionFunctionType = ParametersOfFunction = ReferenceFrequency OutputFileName InputFileOrDirectoryName
to execute map. If the output directory does not exist, an error will occur. Meanings and formats of arguments are as follows:
- CoordinateSystem RADEC, GLGB, or XY
- Projection GLS or ARC
- ProjectCenterPositionX format is hh:mm:ss.ss (RADEC) or dd:mm:ss.ss (GLGB, XY). if you specify DEFAULT , source position (written in the source table) is used
- ProjectCenterPositionY dd:mm:ss.ss or DEFAULT
- BLCPositionX hh:mm:ss.ss (RADEC), dd:mm:ss.ss (GLGB), ssss.ss (XY)
- BLCPositionY dd:mm:ss.ss (RADEC, GLGB), ssss.ss (XY)
- TRCPositionX hh:mm:ss.ss (RADEC), dd:mm:ss.ss (GLGB), ssss.ss (XY)
- TRCPositionY dd:mm:ss.ss (RADEC, GLGB), ssss.ss (XY)
- ZaxisType v (velocity) or f (frequency)
- ZaxisStartPoint The unit is km/s (velocity) or Hz (frequency)
- ZaxisEndPoint The unit is km/s (velocity), Hz (frequency)
- XaxisGridNumber If you specify 0, XaxisGridSpacing takes priority
- YaxisGridNumber If you specify 0, YaxisGridSpacing takes priority
- XaxisGridSpacing The unit is arcsec. If you specify 0, XaxisGridNumber takes priority
- YaxisGridSpacing The unit is arcsec. If you specify 0, YaxisGridNumber takes priority
- ZaxisGridNumber If you specify 0, ZaxisGridSpacing takes priority
- ZaxisGridSpacing The unit is km/s (velocity) or Hz (frequency). If you specify 0, ZaxisGridNumber takes priority
- SearchingRadius The unit is arcsec. Spectra which are closer to the grid point than this radius are convolved
- MinumumNumberOfData If the number of convolved spectra is smaller than this number, the map is blanked at the grid
- ConvolutionFunctionType BG (Bessel*Gauss), SG (Sinc*Gauss), G (Gaussian), PB (Pillbox) or SF (Spheroidal function)
- ParametersOfFunction If multiple parameters are necessary, they are separated by a space.
The "equal" symbols and parameters should also be separated by a space with each other
- BG, SG Parameters a and b.
The unit is arcsec (not pixels).
Set
a=GridSpacing
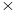 1.55
/
1.55
/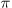 ,
b=GridSpacing
,
b=GridSpacing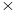 2.52
2.52 - G Parameter a. The unit is arcsec (not pixels). Set a=GridSpacing
- PB Specify 0 as a dummy, though Pillbox function does not have any parameters
- SF
Schwab's parameters m and
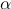
- BG, SG Parameters a and b.
The unit is arcsec (not pixels).
Set
a=GridSpacing
- ReferenceFrequency The unit is Hz. If you specify 0, the value written in the data header is used
- OutputFileName Specify output file name
- InputFileOrDirectoryName Specify input files and/or directories. Multiple files/directories can be specified, and wild-cards can be used (note: the length of the command line should be less than 10240 characters after expansion of wild-cards)
(e.g.) otf_map RADEC GLS DEFAULT DEFAULT 05:35:00.0 -05:40:00.0 5:25:00.0 -05:10:00.0 v -100.0 100.0 0 0 8.0 8.0 0 1.0 24.0 1 BG = 3.947 20.16 = 0 /home/a0011hg/proj0/map/klotf.fits /home/a0011hg/proj0/otfdata/split/klotf.OriKL.*.A01.base /home/a0011hg/proj0/otfdata/split/directoryOfBaselinedFiles